2026
Evolutionary conditioning enables guided generation of functionally diverse enhancers. Duncan AG, Consens ME, Crawford L, Mitchell JA, Moses AM, Yang KK, & Lu AX. bioRxiv 2026 Apr. doi: https://doi.org/10.64898/2026.04.13.718170
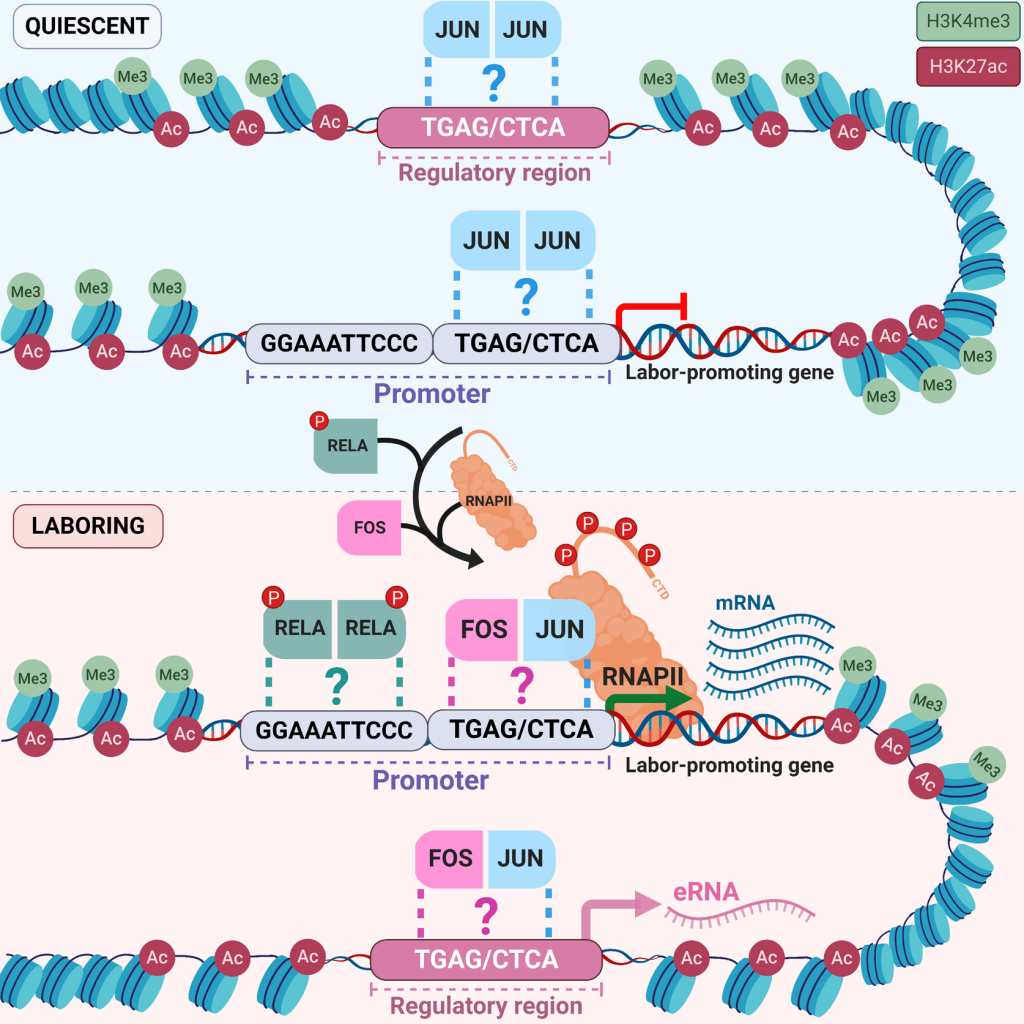
Temporal chromatin and transcriptome dynamics driven by JUND and progesterone receptor binding in the pregnant mouse myometrium. Khader N, Duncan A, Dorogin A, Shynlova O, Mitchell JA. bioRxiv. 2026 Feb. doi: https://doi.org/10.64898/2026.02.25.707818
2025
JUND plays a genome-wide role in the quiescent to contractile switch in the pregnant human myometrium. Khader N, Dorogin A, Shynlova O, Mitchell JA. PLoS Genet. 2025 Jun. doi: https://doi.org/10.1371/journal.pgen.1011261
Lado-Fernández, P., Vilas, J. M., Fernandes, T., Carneiro, C., Da Silva-Álvarez, S., Estévez-Souto, V., Pedrosa, P., González-Barcia, M., Abatti, L. E., Mitchell, J. A., Rivas, C., Moreno-Bueno, G., Vidal, A., & Collado, M. Transcriptional repression of SOX2 by p53 in cancer cells regulates cell identity and migration. Int J Cancer. Published online June 2, 2025 https://doi.org/10.1002/ijc.35490
Abatti L, Gillespie Z, Lado-Fernández P, Collado M, Mitchell JA. A role for NFIB in SOX2 downregulation and epigenome accessibility changes due to long-term estrogen treatment of breast cancer epithelial cells. Biochem Cell Biol. 2025;103:1-14. https://doi.org/10.1139/bcb-2024-0287
A Sox2 enhancer cluster regulates region-specific neural fates from mouse embryonic stem cells. Tobias IC, Moorthy SD, Shchuka VM, Langroudi L, Cherednychenko M, Gillespie ZE, Duncan AG, Tian R, Gajewska NA, Di Roberto RB, Mitchell JA. G3 Genes|Genomes|Genetics. 2025 Apr. doi: 10.1093/g3journal/jkaf012
2024
SOX4 exerts contrasting regulatory effects on labor-associated gene promoters in myometrial cells. Khader N, Shchuka VM, Dorogin A, Shynlova O, Mitchell JA. PLoS One. 2024 Apr 18;19(4):e0297847. doi: 10.1371/journal.pone.0297847.
THE CELLS ARE ALL-RIGHT: Regulation of the Lefty genes by separate enhancers in mouse embryonic stem cells. Taylor T, Zhu HV, Moorthy SD, Khader N, Mitchell JA. https://doi.org/10.1101/2024.06.14.598870
Platania A, Erb C, Barbieri M, Molcrette B, Grandgirard E, de Kort MAC, Pomp W, Meaburn K, Taylor T, Shchuka VM, Kocanova S, Nazarova M, Oliveira GM, Mitchell JA, Soutoglou E, Lenstra TL, Molina N, Papantonis A, Bystricky K, Sexton T. Transcription processes compete with loop extrusion to homogenize promoter and enhancer dynamics. Sci Adv. 2024 Dec 13;10(50):eadq0987. https://doi.org/10.1126/sciadv.adq0987
2023
MYB and ELF3 differentially modulate labor-inducing gene expression in myometrial cells. Shchuka VM, Khader N, Dorogin A, Shynlova O, Mitchell JA. PLoS One. 2023 Jan 3;18(1):e0271081. https://doi.org/10.1371/journal.pone.0271081
Epigenetic misactivation of a distal developmental enhancer cluster drives SOX2 overexpression in breast and lung cancer. Abatti LE, Lado-Fernández P, Huynh L, Collado M, Hoffman MM, Mitchell JA. ucleic Acids Research, Volume 51, Issue 19, 27 October 2023, Pages 10109-10131, https://doi.org/10.1093/nar/gkad734
2022
Histone macroH2A1 is a stronger regulator of hippocampal transcription and memory than macroH2A2 in mice. Singh G, Stefanelli G, Narkaj K, Brimble MA, Creighton SD, McLean TAB, Hall M, Mitchnick KA, Zakaria J, Phung T, Reda A, Leonetti AM, Monks A, Ianov L, Winters BD, Walters BJ, Davidoff AM, Mitchell JA, Zovkic IB. Commun Biol. 2022 May 19;5(1):482. doi: 10.1038/s42003-022-03435-4
Transcriptional regulation and chromatin architecture maintenance are decoupled modular functions at the Sox2 locus. Taylor T, Sikorska N, Shchuka VM, Chahar S, Ji C, Macpherson NN, Moorthy SD, de Kort MAC, Mullany S, Khader N, Gillespie ZE, Langroudi L, Tobias IC, Lenstra TL, Mitchell JA, Sexton T. Genes Dev. 2022 Jun 1;36(11-12):699-717. doi: https://doi.org/10.1101/gad.349489.122
2021
Testing the super-enhancer concept. Blobel GA, Higgs DR, Mitchell JA, Notani D, Young RA. doi: https://doi.org/10.1038/s41576-021-00398-w
Transcriptional Control of Parturition: Insights from Gene Regulation Studies in the Myometrium. Khader N, Shchuka VM, Shynlova O, Mitchell JA. doi: https://doi.org/10.1093/molehr/gaab024
A flexible repertoire of transcription factor binding sites and diversity threshold determines enhancer activity in embryonic stem cells. Singh G, Mullany S, Moorthy SD, Zhang R, Mehdi T, Tian R, Duncan AG, Moses AM, Mitchell JA. Genome Res. 2021 Apr;31(4):564-575. doi: https://doi.org/10.1101/gr.272468.120
2020
Transcriptional enhancers: from prediction to functional assessment on a genome-wide scale. Tobias IC, Abatti LE, Moorthy SD, Mullany S, Taylor T, Khader N, Filice MA, Mitchell JA.Genome. 2020 Sep 22. doi: 10.1139/gen-2020-0104. Online ahead of print.PMID: 32961076
Liang M, Soomro AU, Tasneem S, Abatti LE, Alizada A, Yuan X, Uusküla-Reimand L, Antounians L, Alvi SA, Paterson AD, Rivard GE, Scott IC, Mitchell JA, Hayward CPM, Wilson MD Enhancer-gene rewiring in the pathogenesis of Quebec Platelet Disorder. Blood blood.2020005394. doi: https://doi.org/10.1182/blood.2020005394
Shchuka VM, Abatti LE, Hou H, Khader N, Dorogin A, Wilson MD, Shynlova O, Mitchell JA. The pregnant myometrium is epigenetically activated at contractility-driving gene loci prior to the onset of labor in mice. PLoS Biol 18(7): e3000710. https://doi.org/10.1371/journal.pbio.3000710
2019
Belak ZR, Pickering JA, Gillespie ZE, Audette GF, Eramian M, Mitchell JA, Bridger JM, Kusalik A, Eskiw CH Genes responsive to rapamycin and serum deprivation are clustered on chromosomes and undergo re-organization within local chromatin environments. Biochem Cell Biol, Sept 3, 2019
Dhaliwal NK, Abatti L, Mitchell JA KLF4 protein stability regulated by interaction with pluripotency transcription factors overrides transcriptional control Genes Dev, 2019;33(1): 1-14
Mitchell JA Pluripotency on Lockdown after Deletion of Three Transcription Regulators Preview in Cell Stem Cell 2019;24(5): 681-683
TF Mehdi, G Singh, JA Mitchell, AM Moses. Variational infinite heterogeneous mixture model for semi-supervised clustering of heart enhancers. Bioinformatics 35 (18), 3232-3239
2018
Dhaliwal NK, Miri K, Davidson S, El Jarkass HT, Mitchell JA KLF4 Nuclear Export Requires ERK Activation and Initiates Exit from Naive Pluripotency Stem Cell Reports 2018;10(4): 1308-1323
2017
Moorthy SD, Davidson S, Shchuka VM, Singh G, Malek-Gilani N, Langroudi L, Martchenko A, So V, Macpherson NN and Mitchell JA. Enhancers and super-enhancers have an equivalent regulatory role in embryonic stem cells through regulation of single or multiple genes. Genome Res. 2017;27: 246-258.
2016
Moorthy SD, Mitchell JA. Generating CRISPR/Cas9 Mediated Monoallelic Deletions to Study Enhancer Function in Mouse Embryonic Stem Cells J Vis Exp 2016, (110), e53552
Dhaliwal NK, Mitchell JA. Nuclear RNA Isolation and Sequencing. Methods Mol Biol. 2016;1402:63-71.
2015
Gillespie ZE, MacKay K, Sander M, Trost B, Dawicki W, Wickramarathna A, Gordon J, Eramian M, Kill IR, Bridger JM, Kusalik A, Mitchell JA, Eskiw CH. Rapamycin reduces fibroblast proliferation without causing quiescence and induces STAT5A/B-mediated cytokine production. Nucleus 2015, 6(6):490-506.
Trost B, Moir CA, Gillespie ZE, Kusalik A, Mitchell JA, Eskiw CH. Concordance between RNA-sequencing data and DNA microarray data in transcriptome analysis of proliferative and quiescent fibroblasts. R Soc Open Sci. 2015 Sep 30;2(9):150402
VM Shchuka, N Malek-Gilani, G Singh, L Langroudi, NK Dhaliwal, SD Moorthy, S Davidson, NN Macpherson and JA Mitchell. Chromatin Dynamics in Lineage Commitment and Cellular Reprogramming. Genes 2015, 6(3), 641-661.
S Schoenfelder, M Furlan-Magaril, B Mifsud, F Tavares-Cadete, R Sugar, BM Javierre, T Nagano, Y Katsman, M Sakthidevi, SW Wingett, E Dimitrova, A Dimond, LB Edelman, S Elderkin, K Tabbada, E Darbo, S Andrews, Herman B, A Higgs, E LeProust, CS Osborne, JA Mitchell, NM Luscombe and P Fraser. The pluripotent regulatory circuitry connecting promoters to their long-range interacting elements. Genome Res. April 2015 25: 582-597.
2014
HY Zhou, Y Katsman, NK Dhaliwal, S Davidson, NN Macpherson, M Sakthidevi, F Collura,and JA Mitchell A Sox2 distal enhancer cluster regulates embryonic stem cell differentiation potential. Genes Dev, 2014, 28 (24): 2699-2711.
2013
S Davidson, NN Macpherson, and JA Mitchell. Nuclear organisation of RNA polymerase II transcription. Biochem. Cell Biol. 91: 1-9 (2013)
2012
A Bhattacharya, CY Chen, S Ho and JA Mitchell. Upstream distal regulatory elements contact the Lmo2 promoter in mouse erythroid cells. PLoS ONE, 2012, 7(12): e52880
T Sexton, S Kurukuti, JA Mitchell, D Umlauf, T Nagano and P Fraser. Sensitive detection of chromatin co-associations using enhanced chromosome conformation capture on chip. Nature Protocols, 2012, 7:1335-1350.
JA Mitchell, I Clay, D Umlauf, CY Chen, CA Moir, CH Eskiw, S Schoenfelder, L Chakalova, T Nanano, and P Fraser. Nuclear RNA sequencing of the mouse erythroid cell transcriptome. PLoS ONE, 2012, 7(12): e49274
CY Chen, Q Morris and JA Mitchell. Enhancer identification in mouse embryonic stem cells using integrative modelling of chromatin and genomic features. BMC Genomics, 2012, 13:152.
2010
S Schoenfelder, T Sexton, L Chakalova, NF Cope, A Horton, S Andrews, S Kurukuti, JA Mitchell, D Umlauf, DS Dimitrova, CH Eskiw, Y Luo, CL Wei, Y Ruan, JJ Bieker and P Fraser. Preferential associations between co-regulated genes reveal a transcriptional interactome in erythroid cells. Nature Genetics, 2010 Jan;42(1):53-61.
2008
T Nagano, JA Mitchell, LA Sanz, FM Pauler, AC Ferguson-Smith, R Feil and P Fraser. The Air non-coding RNA epigenetically silences transcription by targeting G9a to chromatin. Science. 2008, 322(5908): 1717-20.
JA Mitchell and P Fraser. Transcription factories are nuclear sub-compartments that remain in the absence of transcription. Genes Dev. 2008, 22(1): 20-5.
2007
J Miles, JA Mitchell, L Chakalova, B Goyenechea, CS Osborne, L O’Neill, K Tanimoto, JD Engel, P Fraser. Intergenic Transcription, Cell-Cycle and the Developmentally Regulated Epigenetic Profile of the Human Beta-Globin Locus. PLoS ONE, 2007, 2(7): e630.
CS Osborne, L Chakalova, JA Mitchell, A Horton, AL Wood, DJ Bolland, AE Corcoran and P Fraser. Myc Dynamically and Preferentially Relocates to a Transcription Factory Occupied by Igh. PLoS Biol. 2007 5(8):e192.
2005
L Chakalova, E Debrand, JA Mitchell, CS Osborne, and P Fraser, Replication and Transcription: Shaping the Landscape of the Genome. Nat Rev Genet, 2005, 6(9): 669-77.
JA Mitchell and SJ Lye, Differential Activation of the Connexin 43 Promoter by Dimers of Activator Protein-1 Transcription Factors in Myometrial Cells. Endocrinology, 2005, 146(4): 2048-54.
2004
CS Osborne, L Chakalova, KE Brown, D Carter, A Horton, E Debrand, B Goyenechea, JA Mitchell, S Lopes, W Reik and P Fraser, Active genes dynamically co-localise to shared sites of ongoing transcription. Nat Genet, 2004, 36 (10): 1065-1071.
JA Mitchell, O Shynlova B Lowell Langille and SJ Lye, Mechanical Stretch and Progesterone Differentially Regulate Activator Protein-1 Transcription Factors in Primary Rat Myometrial Smooth Muscle Cells. Am J of Physiol: Endocrinol Metab, 2004, 287: E439-E445.
2002
JA Mitchell, and SJ Lye, Differential Expression of Activator Protein-1 Transcription Factors in Pregnant Rat Myometrium. Biol Reprod, 2002, 67 (1): 240-246.